2. SMART Protocols Description back to ToC
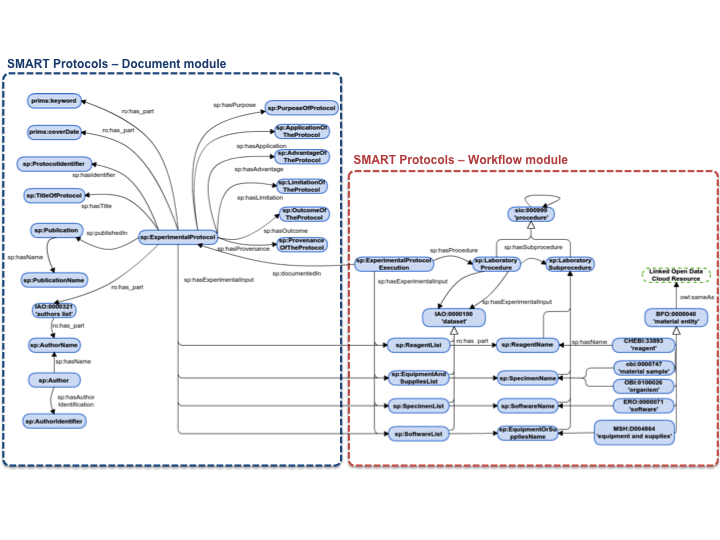
Here we present SMART Protocols (henceforth SP), our ontology-based approach for representing experimental protocols; we aim to "facilitate the semantic representation of experimental protocols". Our ontology reuses BFO; we are also reusing the ontology of relations (RO) to characterize concepts. In addition, each term in the SP ontology is represented with annotation properties imported from OBI Minimal metadata. The classes, properties and individuals are represented by their respective labels to facilitate the readability. The prefix indicates the provenance for each term. Our ontology is organized in two modules. the document module (SP Document Ontology) represent the metadata necesary and suficient for reporting a protocol. The workflow module (SP Workflow Ontology), represent the executable elements of a protocol to be carried out and maintained by humans. A graphical illustration of the ontology modules is presented in Figure 1.
4. References back to ToC
- [BFO]:Robert Arp and Barry Smith. (2008) Function, Role, and Disposition in Basic Formal Ontology. Available from Nature Precedings.
- [OBO_REL]:Barry Smith, Werner Ceusters, Bert Klagges, Jacob Köhler, Anand Kumar, Jane Lomax, Chris Mungall, Fabian Neuhaus, Alan L Rector and Cornelius Rosse. (2005) Relations in biomedical ontologies. Genome Biology, Vol. 6:R46.
Acknowledgements back to ToC
The authors would like to thank Silvio Peroni for developing LODE, a Live OWL Documentation Environment, which is used for representing the Cross Referencing Section of this document and Daniel Garijo for developing Widoco, the program used to create the template used in this documentation.